What if we lowered the barriers to sharing early data, half-formed ideas, and even our uncertainties?
By Lisa Goodrich, Ph.D.
For the past five years, I’ve had the honor of overseeing Hearing Health Foundation’s Hearing Restoration Project (HRP), an extraordinary group of deeply committed, genuinely collaborative scientists.
Scientists become scientists because we are curious. We want to understand how things work. But science often forces us to focus on one very small piece of a very big puzzle.
That’s not a flaw. It’s actually how we build deep expertise. We know our systems, data, and models incredibly well.
Traditionally, we share that knowledge at conferences or through papers. Slowly, piece by piece, we assemble the puzzle.
The HRP was built on a simple but powerful idea: What if we could put the puzzle together faster? What if we didn’t wait for conferences? What if we lowered the barriers to sharing early data, half-formed ideas, and even our uncertainties? What if we truly worked as a team?
With that in mind, the central question the HRP focuses on is this: Why can hair cells regenerate in some species but not in others?
In the human cochlea, when a hair cell dies, it’s gone forever. That’s why hearing loss is permanent for millions of people.
But in fish and birds, the story is very different. Those species can lose many—or even all—of their hair cells, and they grow back. Function returns. Hearing is restored.
So what’s the difference?
In regenerative species, supporting cells, which are the neighboring cells in the sensory epithelium, can detect when a hair cell is lost. Then, remarkably, they can either divide to produce new hair cells or directly convert themselves into hair cells. It’s a natural repair system.
If it were simple, we would just compare fish and humans, identify the difference, and flip the switch. But biology is never that simple. Each species has its own genetic programs, regulatory systems, and molecular context.
This is where HRP comes in. We are 14 principal investigators, each an expert in a specific species or system. Individually, each of us holds a piece of the puzzle. But the HRP is not just a funding mechanism. It’s not a loose network. It’s a true working consortium.
We meet constantly. We share data early. We challenge one another’s interpretations. We formed four working groups focused on specific aspects of regeneration.
We meet as a full consortium every two weeks. Once a year, we gather and decide together what our priorities are and who will do what.
We are guided by HHF’s Scientific Advisory Board with expertise not only in the inner ear, but in regeneration across systems.
From day one, collaboration has not been a slogan. It has been our operating principle.
Scientifically, we believe that if we can understand the molecular changes that allow a supporting cell to become a hair cell in regenerative species, we can identify entry points for regeneration in mammals.
To do that, we generate and analyze large transcriptomic datasets. In simple terms, we look at which genes are turned on and off in hair cells and supporting cells across many species. We want to understand the gene programs that define a hair cell—and the programs that allow plasticity.
But cross-species analysis is hard. First, gene names differ between species. A gene in humans may have multiple related versions, called orthologs, in mice or fish. So we created a unified system. For every human gene, we identified all possible corresponding genes in other species.
Instead of guessing which one matches best, we included them all and let the data guide us.
Second, datasets were generated in different labs, using different methods. That means we can’t simply compare raw expression counts.
So instead, we used a statistical measure called AUC, which stands for “area under the receiver operating curve.” In practical terms, AUC tells us how well a gene distinguishes one cell type from another. An AUC close to 1 means a gene is highly specific for hair cells. An AUC of 0.5 means it’s not specific at all.
For example, the gene otoferlin is a classic hair cell gene. Across datasets and species, its AUC values approach 1 in hair cells. That tells us our approach works.
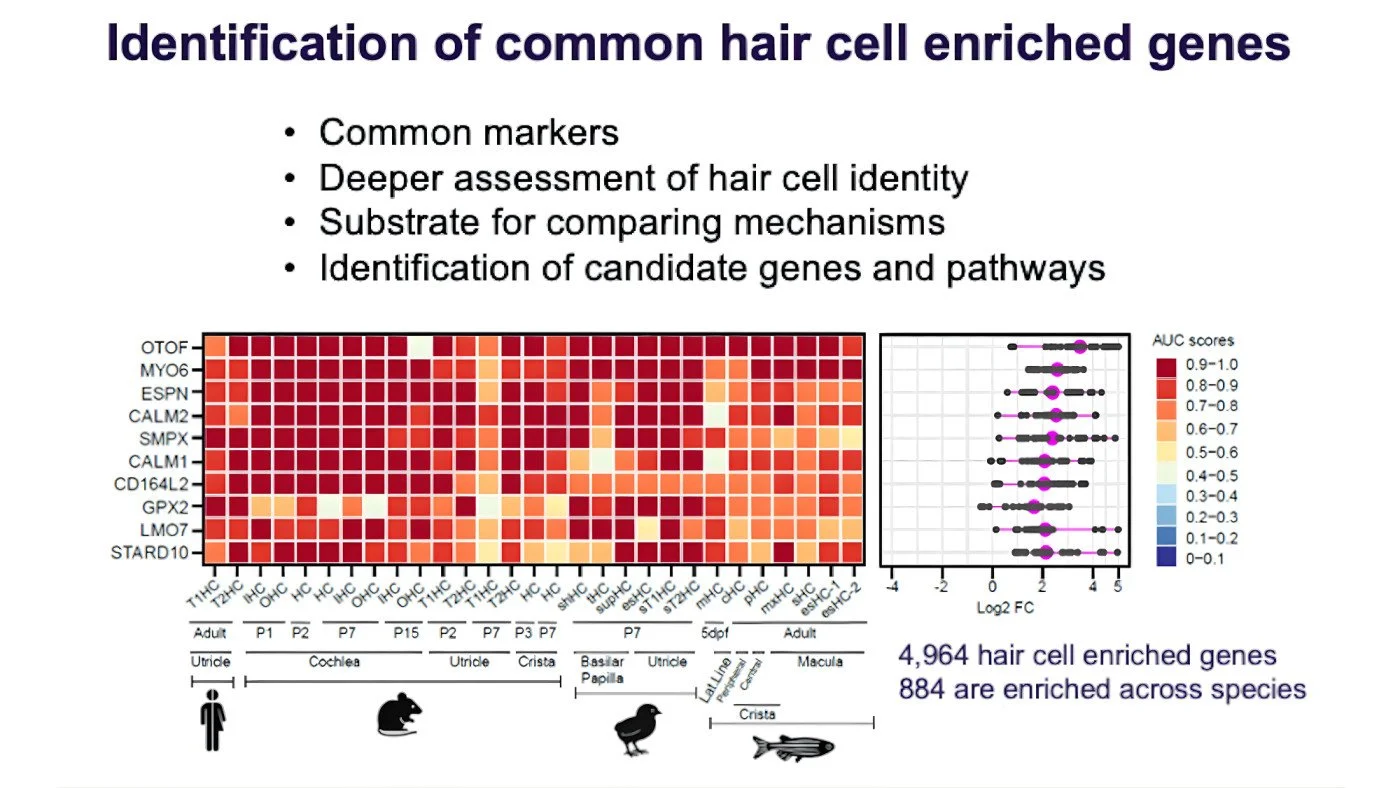
We applied this method genome-wide, across 29 hair cell types from multiple species. We identified nearly 5,000 genes enriched in at least one hair cell type. Then, using unbiased machine learning, we identified a core set of 884 genes consistently enriched across species.
Each red square represents the degree of enrichment relative to supporting cells—the AUC—for the gene name (on the left) in each of 23 hair cell types (listed left to right). The deeper the red, the more enriched. The HRP identified 4,964 hair cell enriched genes and a core set of 884 genes that are consistently enriched in hair cells across species. Credit: The Hearing Restoration Project
This core set is incredibly valuable. First, it gives us common markers, or shared molecular identifiers of hair cells across species. But more importantly, it gives us depth. Turning on one gene like myosin 6 is not enough to create a functional hair cell. Regeneration requires activating an entire gene program. These 884 genes give us a framework for understanding that full identity.
They also allow us to ask deeper mechanistic questions. For example: Why is a particular gene silent in a mouse supporting cell but active in a chicken supporting cell? Is the difference epigenetic? Are regulatory regions locked down in mammals? These are the kinds of questions we can now pursue systematically.
All of this depends on access to shared data. That’s why HRP was an early supporter of gEAR, the gene Expression Analysis Resource.
We needed a common platform where we could upload, explore, and compare transcriptomic datasets.
The gEAR platform is now used worldwide.
Through the gEAR, anyone can explore our analyses. Scientists can enter a gene of interest and immediately see its enrichment patterns across hair cell types and species. They can examine fold changes, AUC values, and even return to the original datasets.
We also built a tool called the HRP Hair Cell Gene Explorer. It allows users to customize analyses. Researchers can select specific hair cell types—up to 43 different ones—and generate tailored plots. They can then define their own criteria: for example, “Show me genes highly enriched in human utricular hair cells but not in mouse cochlear hair cells.”
The system will generate a gene list that meets the researchers’ thresholds. It can be downloaded to use in their own experiments.
Importantly, our cross-species ortholog mapping prevents misleading conclusions. If a human gene corresponds to two mouse genes, the tool shows both. We can see which ortholog is truly expressed, rather than accidentally concluding a gene is absent.
I can’t emphasize enough how this entire infrastructure exists because we chose to work collaboratively. Our Zoom meetings are lively. We debate. We refine. We bring trainees into the meetings. (I love when I finally meet in person the students and postdocs I’ve been seeing on screen for months!)
We are building not just datasets but also a community.
And I want to emphasize another point: Although our focus has been hair cell regeneration, the tools we’ve built can be used much more broadly. We are the Hearing Restoration Project—not just the “Hair Cell Restoration Project.” We have not yet expanded deeply into neurons or ganglion cells, but the framework we’ve created could absolutely support that work. The nomenclature system, cross-species comparisons, and enrichment tools—all of it is adaptable.
What excites me most is that this isn’t just about what our consortium can do. It’s about what all hearing and balance researchers can now do with these tools.
If we want to solve hair cell regeneration, it will not happen in isolated labs working in parallel. It will happen through shared data, shared frameworks, shared questions. That was our founding belief, that lowering barriers to collaboration would accelerate discovery.
I am grateful for our Scientific Advisory Board, our investigators, and especially our trainees whose work drives so much of this progress. We are still assembling the puzzle. But we are assembling it together, and that makes all the difference.
This is adapted from the presentation Lisa Goodrich, Ph.D., made as part of a panel about collaboration at the Association of Research in Otolaryngology MidWinter Meeting in February 2026. The scientific director of the HRP, Goodrich is a professor of neurobiology at Harvard Medical School.






Veterans, when compared to the general public, have had more occupational exposure to loud noises, such as gunfire and explosions.